This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
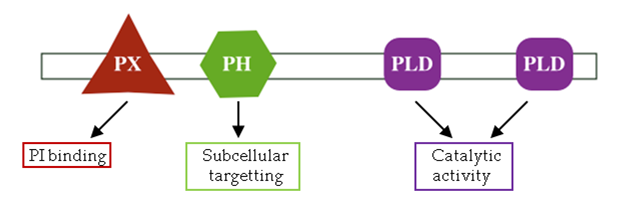
Protein domainsA protein domain is a structure within a protein that has been independently folded, may or may not have a characterized biological function, and is capable of existing outside of the protein. Bioinformatic tools exist that can determine what protein domains are in a protein given the proteins amino acid sequence. After a protein domain is determined from the sequence, the function, structure, and role of the protein at the cellular and organismal levels is often revealed. Unless science has yet to characterize the function of the domain in question.
|
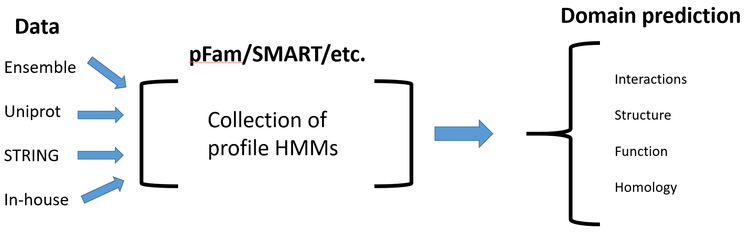
Protein domain databases
Are the bioinformatic tools used for protein domain analysis, they are often free and online. Here are the under-the-hood mechanics of how these databases work.
Data - protein sequences and domain information from outside sources
Database (pFam/SMART/etc.) - Perform multiple sequence alignments and represent these multiple sequence alignments as a profile-hidden Markov model.
Multiple sequence alignment - Sequences of DNA or proteins are stacked on top of each other and each amino acid or nucleotide in the sequence is compared to all the others.
Profile - hidden Markov model (profile HMM) - Statistical model that offers a succinct representation of this multiple sequence alignment. After submitting a sequence to a database the sequence is searched against these profile-HMMs.
Domain prediction - Output from the database, the predicted protein domain(s) and other info about the protein domain(s) and entire protein.
Are the bioinformatic tools used for protein domain analysis, they are often free and online. Here are the under-the-hood mechanics of how these databases work.
Data - protein sequences and domain information from outside sources
Database (pFam/SMART/etc.) - Perform multiple sequence alignments and represent these multiple sequence alignments as a profile-hidden Markov model.
Multiple sequence alignment - Sequences of DNA or proteins are stacked on top of each other and each amino acid or nucleotide in the sequence is compared to all the others.
Profile - hidden Markov model (profile HMM) - Statistical model that offers a succinct representation of this multiple sequence alignment. After submitting a sequence to a database the sequence is searched against these profile-HMMs.
Domain prediction - Output from the database, the predicted protein domain(s) and other info about the protein domain(s) and entire protein.
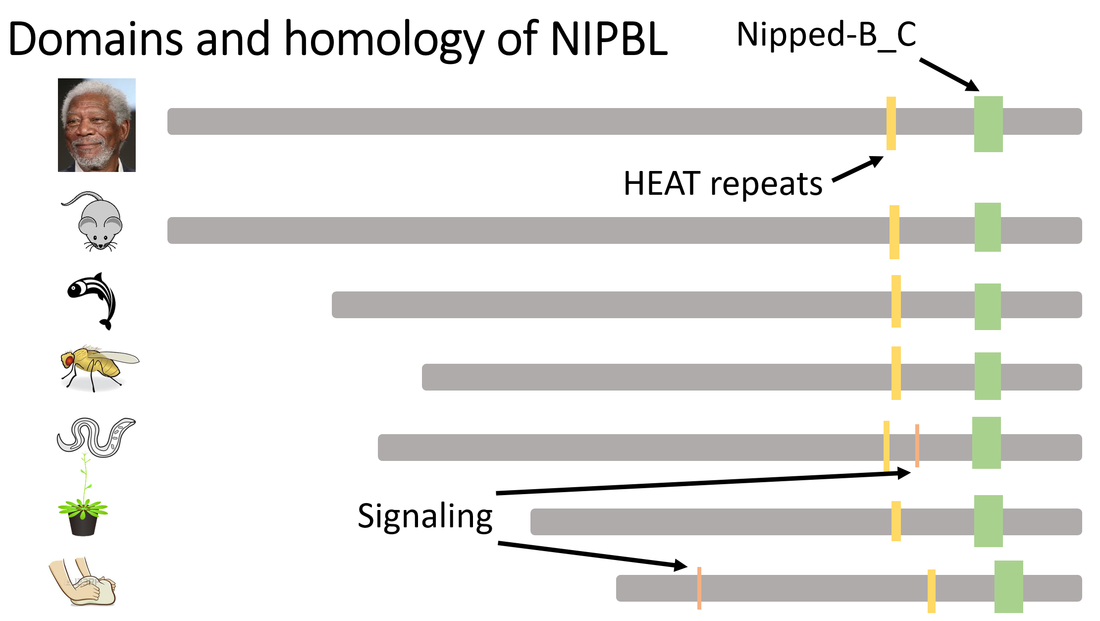
NIPBL protein domain analysis
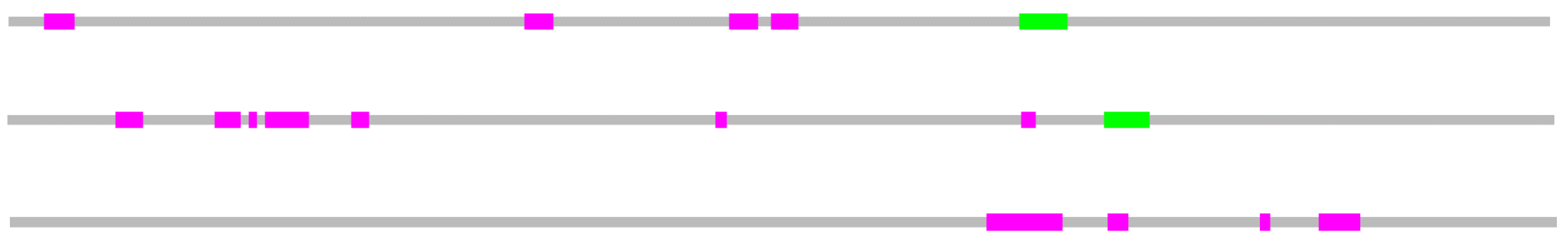
Results of SMART using accession number NP_597677.2
Results of pFam using accession number NP_597677.2
NIPBL has two domains that are well conserved through evolutionary time: the Nipped-B_C domain and the HEAT repeats. Two signaling domains of unknown function also appear in S. cerevisiea and C. elegans. Interestingly both the HEAT repeats and the Nipped-B_C domain are on the less conserved N' terminal half of the protein. The fact that the conserved carboxy terminal half of the protein is well conserved but has no annotated domains is quite interesting. A possible avenue of NIPBL research could be to annotate this conserved region and possibly uncover a previously unknown protein domain.
References:
1.) Toll-Riera, M., Rado-Trilla, N., Martys, F., & Alba, M. (2011). Role of Low-Complexity Sequences in the Formation of Novel Protein Coding Sequences [Abstract]. Oxford Academic: Molecular Biology and Evolution,29(3), 883-886. Retrieved March 15, 2018, from https://academic.oup.com/mbe/article/29/3/883/1007217.
2.) Family: Cohesin_HEAT (PF12765). (n.d.). Retrieved March 17, 2018, from http://pfam.xfam.org/family/PF12765.6
3.) Family: Nipped-B_C (PF12830). (n.d.). Retrieved March 15, 2018, from http://pfam.xfam.org/family/PF12830.6
4.) European Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. (n.d.). European Bioinformatics Institute. Retrieved March 17, 2018, from https://www.uniprot.org/help/coiled
1.) Toll-Riera, M., Rado-Trilla, N., Martys, F., & Alba, M. (2011). Role of Low-Complexity Sequences in the Formation of Novel Protein Coding Sequences [Abstract]. Oxford Academic: Molecular Biology and Evolution,29(3), 883-886. Retrieved March 15, 2018, from https://academic.oup.com/mbe/article/29/3/883/1007217.
2.) Family: Cohesin_HEAT (PF12765). (n.d.). Retrieved March 17, 2018, from http://pfam.xfam.org/family/PF12765.6
3.) Family: Nipped-B_C (PF12830). (n.d.). Retrieved March 15, 2018, from http://pfam.xfam.org/family/PF12830.6
4.) European Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. (n.d.). European Bioinformatics Institute. Retrieved March 17, 2018, from https://www.uniprot.org/help/coiled