Homology
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Sequence homology is the hypothesis that two organisms have similar sequences for a particular gene or protein because they share a common ancestor [1]. Scientist form these hypotheses based off of the percent identity (exact amino or nucleic acid match), which regions of the sequence have the greatest identity, and if the protein has a similar function.
|
Homo sapiens:
Accession: NP_597677 Length: 2804 amino acids [Caenorhabditis elegans] Nematode:
Accession: WBGene00004166 Length: 2203 amino acids Identity: 17.08 % pqn-85 [Danio rerio] Zebrafish:
Accession: ENSDARG00000061052 Length: 2381 amino acids Identity: 83.0 % nipblb |
[Arabidopsis thaliana] Thale Cress:
Accession: EMB2773 Length: 1846 amino acids Identity: 25.0 % AtSCC2 [Nannospalax galili] Upper Galilee mountains blind mole rat:
Accession: ENSNGAG00000018174 Length: 2798 amino acids Identity: 97.4 % |
[Pteropus vampyrus] Megabat:
Accession: ENSPVAG00000001535 Length: 2800 amino acids Identity: 93.62 % [Dictyostelium discoideum] Slimemold:
Accession: XP_639229.1 Length: 2063 amino acids Identity: 35.0 % |
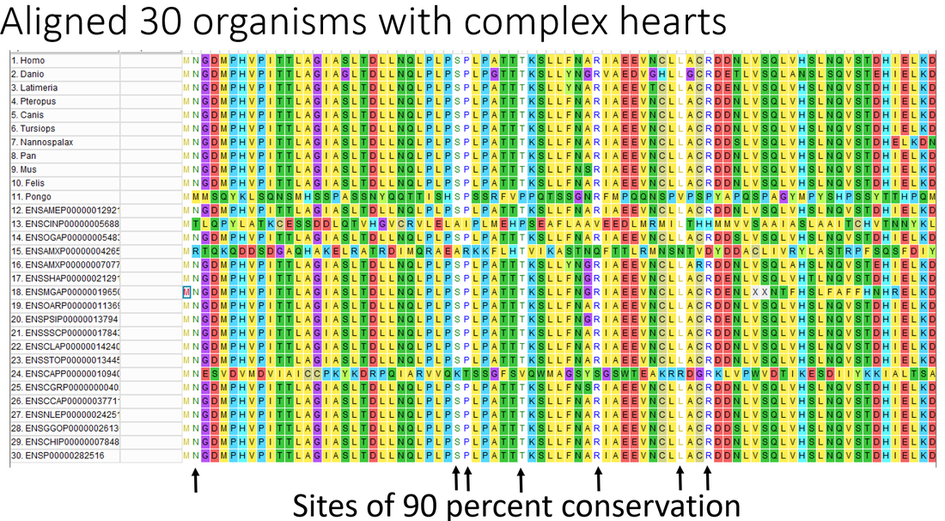
Conservation of carboxy terminus in NIPBL
After collecting the NIPBL sequences for all of the organisms above I manually looked for regions of conservation between organisms. I found that the carboxy terminal of the protein is highly conserved while the N terminal shares little sequence similarity between organisms.
After collecting the NIPBL sequences for all of the organisms above I manually looked for regions of conservation between organisms. I found that the carboxy terminal of the protein is highly conserved while the N terminal shares little sequence similarity between organisms.
1.) Sequence homology. (n.d.). Retrieved April 13, 2018, from https://www.biology-online.org/dictionary/Sequence_homology