Biological networks
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
A biological network is a group of biological entities that is interacting with each other. In our case these entities our small biomolecules (such as DNA, proteins, metabolites, etc) that are either directly binding each other or are regulating their abundances.
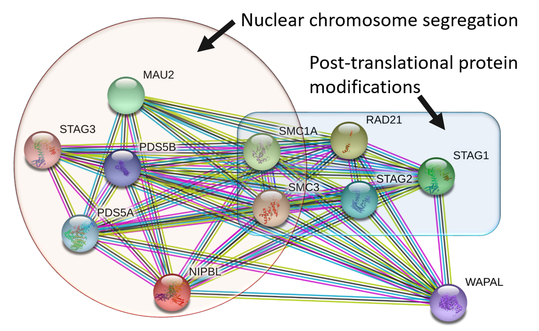
NIPBL interaction network
|
STRING is a free website which visualizes biological networks for a searched protein. To the left is the STRING output for an NIPBL query and my post-search selection of biological process GO terms. Selecting GO terms after a STRING search allows scientist to cluster proteins and potentially identify new roles for proteins of interest.
|
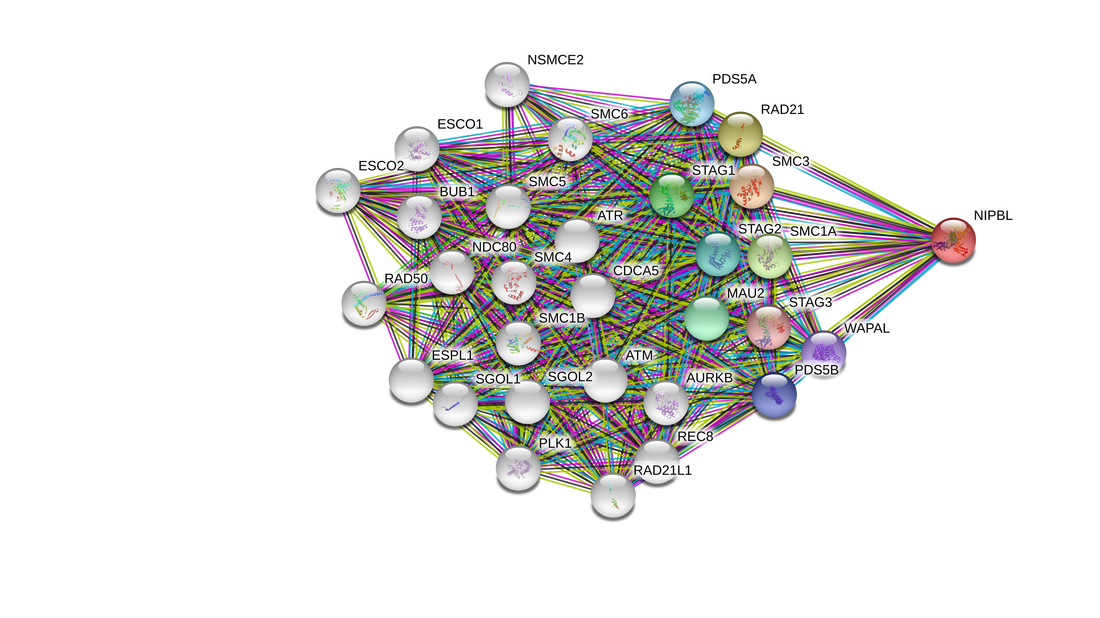
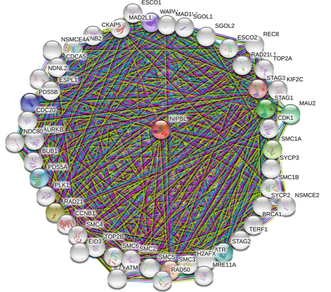
NIPBL interaction network complexity
By selecting the "More" tab after a search on STRING more of the interacting network is displayed. Within a few clicks on the "More" tab the network quickly becomes incomprehensible. Instead of displaying all possible interactions to the user STRING decides what to display based off of scores computed from the observed data and corrects for the probability of randomly observing an interaction [1].
How is STRING data generated?
References
1.) Frequently Asked Questions. (n.d.). Retrieved April 13, 2018, from https://string-db.org/cgi/info.pl?sessionId=EBTgoQE7BnXf&footer_active_subpage=faqs
1.) Frequently Asked Questions. (n.d.). Retrieved April 13, 2018, from https://string-db.org/cgi/info.pl?sessionId=EBTgoQE7BnXf&footer_active_subpage=faqs