This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Model Organisms
|
Model organisms are a research tool that is simple in concept. Basically; use a non-human organism to study biological phenomena while controlling some component of the experiment (like the organism's DNA, environment, food, etc.). Since model organisms have been in use for such a long time this has allowed us (the human species) to build up appreciable amounts of knowledge about these organisms. In fact, databases exist whose sole purpose is to collect information on model organisms (Zebrafish Database, Mouse Database, etc).
|
Choosing the right model organism
When choosing a model organism a scientist has to consider the ethics, cost, previous studies, human similarity, and generation time that differs between model organisms. A general trend when deciding between model organisms is that as they increase in size the cost, ethical considerations, similarity to humans, and generation time increases. To visualize this scenario imagine the differences that would exist between research in a worm (such as C. elegans) vs. performing research in a chimpanzee.
Zebrafish as a model for CdLS
Zebrafish is a human NIPBL orthologue and known to be a model organism that is good for embryological developmental disorders [3]. CdLS is a multi-organ system birth defect which makes the fact that Zebrafish embryos are “see through” and internal development can be observed as the embryo develops, very enticing [1]. One of the characteristic of CdLS is skeletal abnormalities, which makes the Zebrafish a better model than say, C.elegans or D. Melanogaster, because it has a skeleton. Zebrafish was more preferable than mice because of its chemical screening capabilities, cost, generation time, and the previous work that has been performed in the organism [3, 2].
Modeling CdLS heart defects
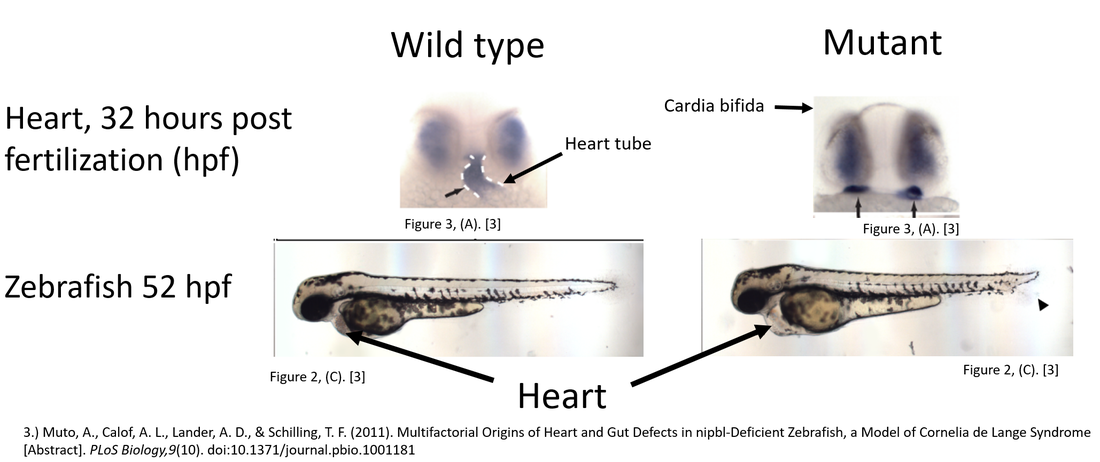
NIPBL is highly expressed in heart and skeletal tissues during development which explains why heart abnormalities are a characteristic of CdLS [4, 5]. Previous studies of NIPBL in zebrafish have revealed what heart specific phenotypes are associated with Zebrafish that have decreased NIPBL expression [1]. The mutant heart phenotype can be seen 32 hours post fertilization (hpf) (pictured below), mutants may have beating hearts but reduced jogging to the left (heart orientation relative to dorsal symmetry is off), may never develop a midline heart tube, and develop cardia bifida (development of two hearts) [1, 6, 7]. At 52 hpf a second observable phenotype emerges: excess fluid in the pericardial cavity [1].
References
1.) Muto, A., Calof, A. L., Lander, A. D., & Schilling, T. F. (2011). Multifactorial Origins of Heart and Gut Defects in nipbl-Deficient Zebrafish, a Model of Cornelia de Lange Syndrome [Abstract]. PLoS Biology,9(10). doi:10.1371/journal.pbio.1001181
2.) Muto, A., & Schilling, T. (2017). Zebrafish as a Model to Study Cohesin and Cohesinopathies [Abstract]. Methods Mol Biol.,177-196. doi:10.1007/978-1-4939-6545-8_11
3.) Zebrafish Development. (n.d.). Retrieved March 15, 2018, from https://embryology.med.unsw.edu.au/embryology/index.php/Zebrafish_Development
4.) Muto, A., Calof, A., Lander, A., & Schilling, T. (2011). Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. PLoS Biol. doi:https://doi.org/10.1371/journal.pbio.10011815.) Strachan, T. (2005). Cornelia de Lange Syndrome and the link between chromosomal function, DNA repair and developmental gene regulation. Current Opinion in Genetics & Development,15(3), 258-264. doi:10.1016/j.gde.2005.04.005
6.) Khodiyar, V. K., Howe, D., Talmud, P. J., Breckenridge, R., & Lovering, R. C. (2013). From zebrafish heart jogging genes to mouse and human orthologs: Using Gene Ontology to investigate mammalian heart development. F1000Research. doi:10.12688/f1000research.2-242.v
7.) (n.d.). Retrieved April 12, 2018, from http://www.informatics.jax.org/vocab/mp_ontology/MP:0004187
1.) Muto, A., Calof, A. L., Lander, A. D., & Schilling, T. F. (2011). Multifactorial Origins of Heart and Gut Defects in nipbl-Deficient Zebrafish, a Model of Cornelia de Lange Syndrome [Abstract]. PLoS Biology,9(10). doi:10.1371/journal.pbio.1001181
2.) Muto, A., & Schilling, T. (2017). Zebrafish as a Model to Study Cohesin and Cohesinopathies [Abstract]. Methods Mol Biol.,177-196. doi:10.1007/978-1-4939-6545-8_11
3.) Zebrafish Development. (n.d.). Retrieved March 15, 2018, from https://embryology.med.unsw.edu.au/embryology/index.php/Zebrafish_Development
4.) Muto, A., Calof, A., Lander, A., & Schilling, T. (2011). Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. PLoS Biol. doi:https://doi.org/10.1371/journal.pbio.10011815.) Strachan, T. (2005). Cornelia de Lange Syndrome and the link between chromosomal function, DNA repair and developmental gene regulation. Current Opinion in Genetics & Development,15(3), 258-264. doi:10.1016/j.gde.2005.04.005
6.) Khodiyar, V. K., Howe, D., Talmud, P. J., Breckenridge, R., & Lovering, R. C. (2013). From zebrafish heart jogging genes to mouse and human orthologs: Using Gene Ontology to investigate mammalian heart development. F1000Research. doi:10.12688/f1000research.2-242.v
7.) (n.d.). Retrieved April 12, 2018, from http://www.informatics.jax.org/vocab/mp_ontology/MP:0004187