Photo : Whole Drosophila sp. (fruit fly) ovary, Dr. Denise Montell : Johns Hopkins School of Medicine
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
This webpage was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
Introductions
Cornelia de Lange Syndrome (CdLS) is a developmental disorder characterized by skeletal abnormalities, hirsutism, mental retardation, and heart defects [1]. CdLS is caused by mutations that affect the cohesin complex, which changes chromatin structure during the processes of DNA repair, transcription, and sister chromatid cohesion [1, 2]. Current evidence suggest that transcriptional dysregulation is the primary cause of CdLS, as CdLS individuals experience transcriptional dysregulation in hundreds of genes and have a functional cohesion process [1, 3]. Fifty-five percent of CdLS cases are caused by mutations in Nipped B-like Protein (NIPBL) which loads cohesin onto DNA [4]. NIPBL expression changes between tissues and is highly expressed in skeletal and heart tissues [5]. Congenital heart defects are present in 30 to 40 percent of CdLS patients and previous studies have confirmed the role of NIPBL in heart development using an NIPBL deficient Zebrafish model [1, 6]. Additionally, 25 percent of CdLS cases are caused by missense mutations and single amino acid deletions in the conserved carboxy-terminal half of NIPBL [5]. It is currently unknown how mutations in these conserved amino acids of NIPBL effect heart development. My primary goal is to determine which amino acids in NIPBL are most critical during heart development and why. Zebrafish is an adequate model for this purpose because it can be used to screen chemical libraries and heart formation can be easily observed during development [6]. I hypothesize that single amino acid mutations in NIPBL will have effects on heart development unique to the amino acid mutated. My long-term goal is to determine the role of NIPBL in mediating heart development.
Aim 1
Determine NIPBL amino acids critical to heart development
Rationale: The amino acids of NIPBL that differ between organisms with complex hearts and organisms with simple hearts may be critical to heart development.
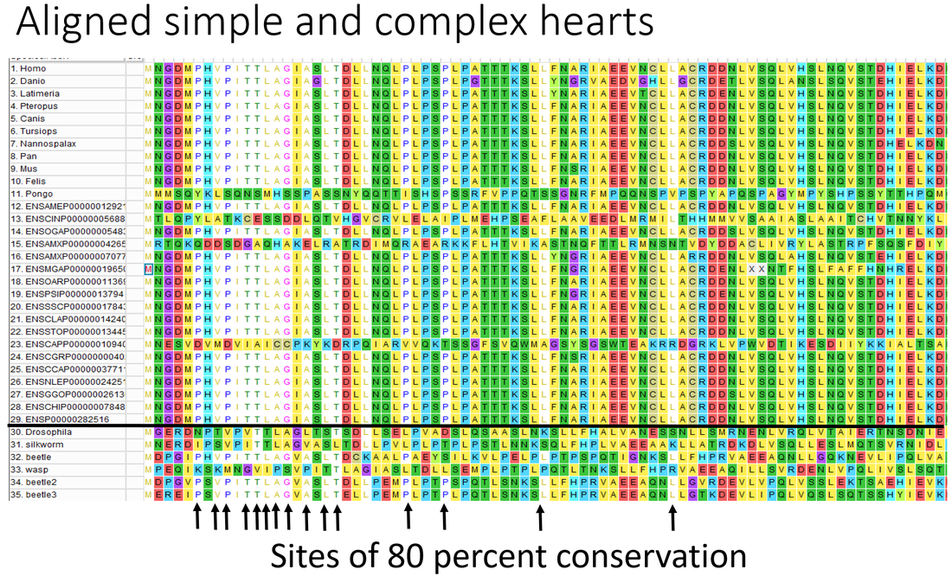
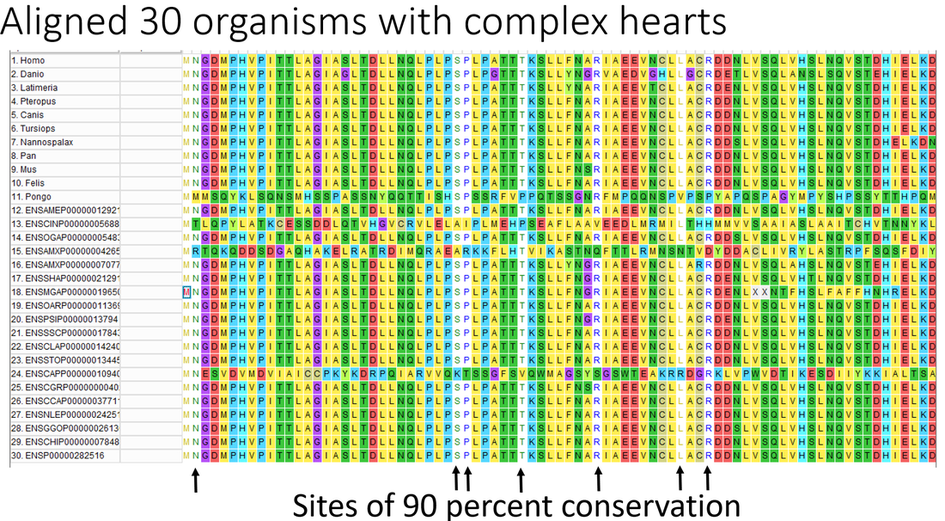
Approach: I used the ClustalW algorithm to align NIPBL sequences of organisms with complex hearts and organisms with simple hearts to determine which amino acids are unique to organisms with complex hearts. First, I performed an alignment between 35 organisms that have both complex and simple hearts (Figure 1). I found 17 amino acids that were moderately conserved between all 35 species. Defining moderate conservation as positions that have an amino acid shared by at least 80 percent of the organisms in the alignment. Then I performed an alignment between 30 organisms with complex hearts and found 7 highly conserved amino acids (Figure 2). Highly conserved sites defined as 90 percent shared amino acid identity.
Rationale: The amino acids of NIPBL that differ between organisms with complex hearts and organisms with simple hearts may be critical to heart development.
Approach: I used the ClustalW algorithm to align NIPBL sequences of organisms with complex hearts and organisms with simple hearts to determine which amino acids are unique to organisms with complex hearts. First, I performed an alignment between 35 organisms that have both complex and simple hearts (Figure 1). I found 17 amino acids that were moderately conserved between all 35 species. Defining moderate conservation as positions that have an amino acid shared by at least 80 percent of the organisms in the alignment. Then I performed an alignment between 30 organisms with complex hearts and found 7 highly conserved amino acids (Figure 2). Highly conserved sites defined as 90 percent shared amino acid identity.
|
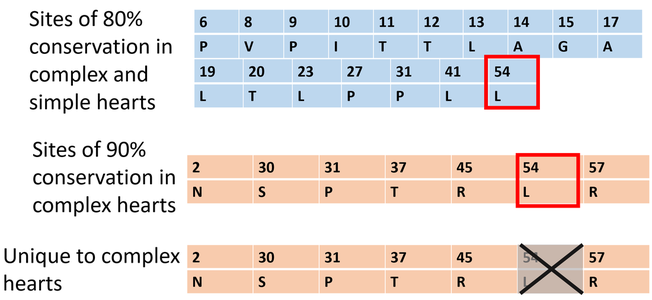
Each column in Figure 4 represents a conserved amino acid; the position is on top and a prefix for the amino acid is below. Lysine at position 54 was moderately conserved in all organisms aligned and also highly conserved in organisms with complex hearts. As its conservation is not specific to organisms with complex hearts it will not be considered critical to complex heart development.
I will then use the CRISPR-Cas system to mutagenize Zebrafish at amino acids "Unique to complex hearts" (Figure 3). |
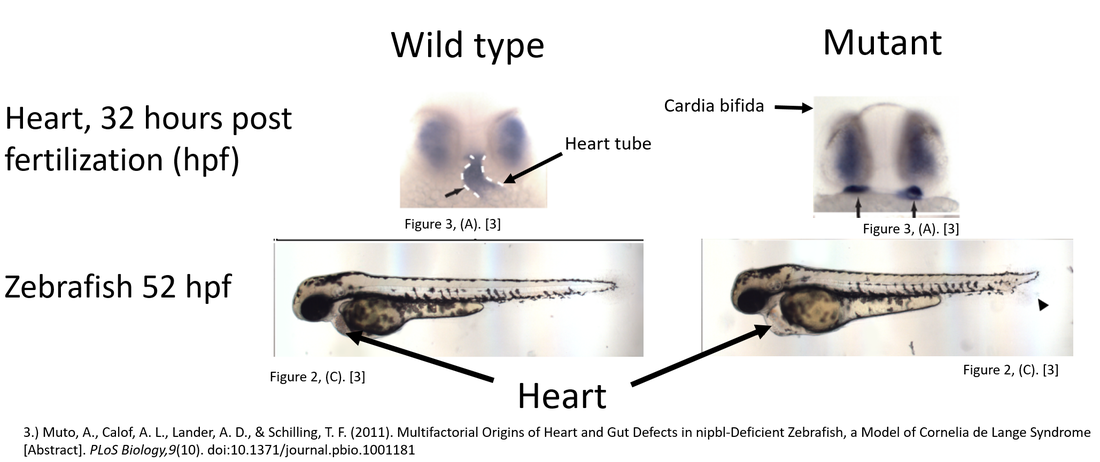
Hypothesis: Zebrafish with mutations in amino acids unique to organisms with complex hearts will exhibit a mutant phenotype. After mutagenisis of the six unique amino acids found in Figure 3 I will observe the Zebrafishes' development. I expect that if the amino acid mutagenized is critical to heart development the mutant's heart morphology will be similar to the heart morphology of an NIPBL deficient Zebrafish (Figure 4). Only amino acids whose mutations resulted in a mutant phenotype will be selected for continued study.
After 32 hours of development the NIPBL deficient Zebrafish has undergone cardia bifida and lacks a midline heart tube [6]. The mutant Zebrafish may display a second phenotype 52 hpf in which it has excess fluid in the pericardial cavity [6].
Aim 2
Search for small molecules with amino acid specificity
Rationale: Small molecules may exist in chemical libraries capable of rescuing heart defects and interacting with NIPBL at specific amino acids.
Approach: I will screen broad and biological function-oriented libraries for small molecules capable of rescuing the NIPBL mutant phenotype.
Rationale: Small molecules may exist in chemical libraries capable of rescuing heart defects and interacting with NIPBL at specific amino acids.
Approach: I will screen broad and biological function-oriented libraries for small molecules capable of rescuing the NIPBL mutant phenotype.
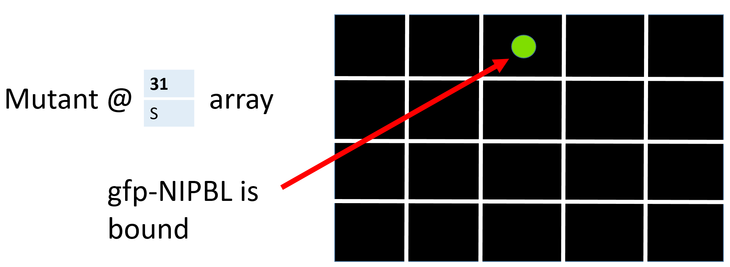
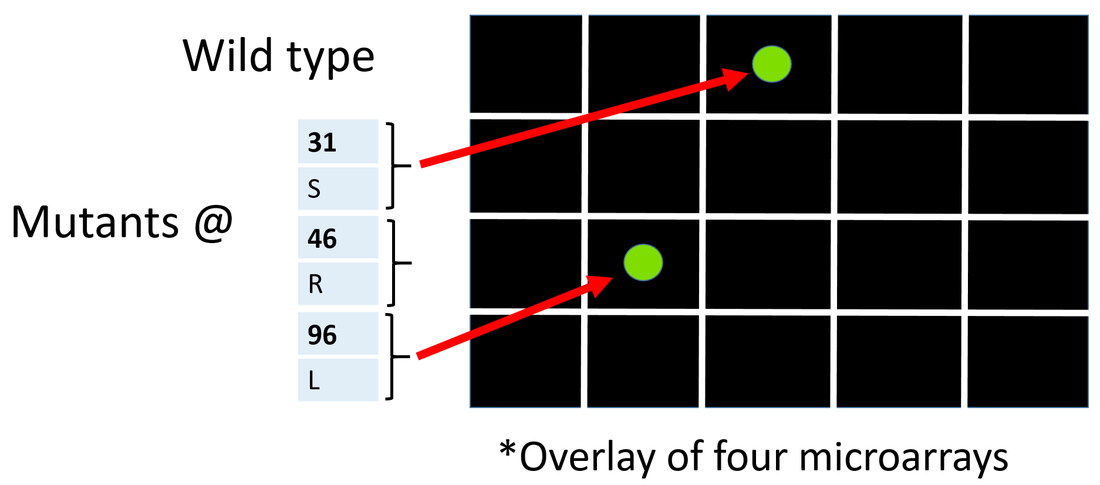
To screen the broad chemical library for NIPBL interacting proteins I will use a small molecule microarray (Figure 5). Using the CRISPR-cas system I will attach green flourescent protein onto the poorly conserved N terminus of each amino acid mutant and wild type NIPBL. Then I will individually wash the different NIPBL fusion proteins over the microarry.
A "hit" in this biochemical screen would be a small molecule that binds to mutagenized NIPBL (Figure 7). Hits from the biochemical screen will then undergo phenotypic screening to see if the "hit" small molecule can rescue the mutant phenotype.

I will also search for potential therapeutics in a biological function-oriented chemical library. Using phenotypic screening I will search The Stem Cell Signaling Compound Library and the Histone Modification Research Compound Library from MedChem Express for potential therapeutics. The Stem Cell Signaling Compound Library was selected because NIPBL had gene ontology (GO) terms for stem cell population maintenance, stem cell development, and stem cell differentiation [7]. The Histone Modification Library was selected because NIPBL loads cohesin onto DNA and cohesin binding is enriched at gene promoters with histone 3 lysine 4 trimethylation [8].
Hypothesis: Potential therapeutics exist in broad chemical libraries capable of rescuing the mutant phenotype via amino acid specific mechanisms. Likewise potential therapeutics may exist in function oriented chemical libraries capable of rescuing the mutant phenotype via non-amino acid specific mechanisms.
Aim 3
Discover amino acid specific interactions
Rationale: Determining which protein-protein interactions of NIPBL change between amino acid mutants will indicate which amino acids are involved when NIPBL binds to other proteins. Subsequent gene ontology (GO) analysis of the discovered interacting partners will indicate how NIPBL directly controls heart development.
Approach: I will attach a TAP-tag onto the less conserved N’ terminus of NIPBL and extract the protein from Zebrafish heart tissue at multiple points of development from WT and each amino acid mutant. NIPBL complexes will then be captured using tandem affinity purification and characterized using mass spectrometry and bioinformatic analysis. Then I will document which interacting partners (proteins) had GO terms for heart development. To validate these partners as proteins critical to heart development I will use the CRISPR-Cas system to knock-out each interacting partner. I will then observe the phenotype of each interacting knock-out.
Rationale: Determining which protein-protein interactions of NIPBL change between amino acid mutants will indicate which amino acids are involved when NIPBL binds to other proteins. Subsequent gene ontology (GO) analysis of the discovered interacting partners will indicate how NIPBL directly controls heart development.
Approach: I will attach a TAP-tag onto the less conserved N’ terminus of NIPBL and extract the protein from Zebrafish heart tissue at multiple points of development from WT and each amino acid mutant. NIPBL complexes will then be captured using tandem affinity purification and characterized using mass spectrometry and bioinformatic analysis. Then I will document which interacting partners (proteins) had GO terms for heart development. To validate these partners as proteins critical to heart development I will use the CRISPR-Cas system to knock-out each interacting partner. I will then observe the phenotype of each interacting knock-out.
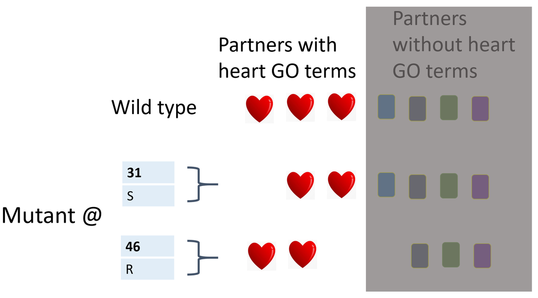
Hypothesis: Interacting proteins that have GO terms for heart development that appear in WT but differ between amino acid mutants are binding partners critical to heart development. Consequently the mutagenized amino acid in an amino acid mutant missing heart GO term partners, is an amino acid which binds these critical proteins (Figure 8). I expect that the CRISPR -Cas knock-out of a binding partner will result in a mutant phenotype.
Future Directions
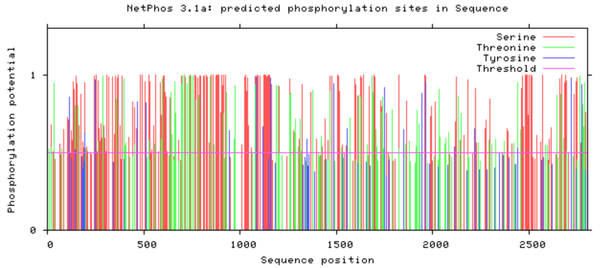
Determining which amino acids in NIPBL are critical to heart development, why they are critical, and how their function can be recapitulated with small molecules following mutation, is an important step towards curing CdLS heart deffects. This research will provide the beginning of an answer to those questions by testing amino acids of likely importance, putting NIPBL into context among its interacting partners, and finding potential drug candidates. Future steps towards fully answering these questions and the larger question of how NIPBL mediates heart development will involve phosphoproteomic studies and continued chemical genetics research. It was predicted in-silico that the conserved carboxy terminal half of NIPBL is heavily phosphorylated (Figure 9) [3]. The combined arguments of amino acid conservation and predicted phosphorylation of conserved beckons for future studies. After this initial drug discovery research is completed I plan to continue chemical genetic research using a higher order model organisms (e.g. mouse) and focused chemical libraries derived from the biochemical "hits" of Aim 2.
References:
1.) Mills, J., Herrera, P., & Kaur, M. et al. (2018). NIPBL /− haploinsufficiency reveals a constellation of transcriptome disruptions in the pluripotent and cardiac states. Scientific Reports. doi:doi:10.1038/s41598-018-19173-9
2.) Remeseiro, S., Cuadrado, A., & Losada, A. (2013). Cohesin in development and disease. Development,140(18), 3715-3718. doi:10.1242/dev.090605
3.) Boudaoud, I., Fournier, É, Baguette, A., & Vallée, M. (2017). Connected Gene Communities Underlie Transcriptional Changes in Cornelia de Lange Syndrome. Genetics,207(1), 139-151. doi:10.1534/genetics.117.202291
4.) Ball, A., Chan, Y., & Yokomori, K. (2014). Mechanisms of cohesin-mediated gene regulation and lessons learned from cohesinopathies. Biochimica Et Biophysica Acta,1839(3), 191-202. doi:https://doi.org/10.1016/j.bbagrm.2013.11.002
5.) Strachan, Tom. “Cornelia De Lange Syndrome and the Link between Chromosomal Function, DNA Repair and Developmental Gene Regulation.” Current Opinion in Genetics & Development, vol. 15, no. 3, 2005, pp. 258–264., doi:10.1016/j.gde.2005.04.005.
6.) Muto, A., Calof, A. L., Lander, A. D., & Schilling, T. F. (2011). Multifactorial Origins of Heart and Gut Defects in nipbl-Deficient Zebrafish, a Model of Cornelia de Lange Syndrome [Abstract]. PLoS Biology,9(10). doi:10.1371/journal.pbio.1001181
7.) STRING Search NIPBL. (n.d.). Retrieved April 13, 2018, from https://string-db.org/
8.) Newkirk, D., Chen, Y., Chien, R., & Zeng, W. (2017). The Effect Of Nipped-B-Like (Nipbl) Haploinsufficiency On Genome-Wide Cohesin Binding And Target Gene Expression: Modeling Cornelia de Lange Syndrome. Clinical Epigenetics,9(89). doi:10.1101/134825
9.) NetPhos 3.1 Server NIPBL prediction. (n.d.). Retrieved April 15, 2018, from http://www.cbs.dtu.dk/services/NetPhos/
10.) Phosphorylation. (2018, May 09). Retrieved from https://en.wikipedia.org/wiki/Phosphorylation
1.) Mills, J., Herrera, P., & Kaur, M. et al. (2018). NIPBL /− haploinsufficiency reveals a constellation of transcriptome disruptions in the pluripotent and cardiac states. Scientific Reports. doi:doi:10.1038/s41598-018-19173-9
2.) Remeseiro, S., Cuadrado, A., & Losada, A. (2013). Cohesin in development and disease. Development,140(18), 3715-3718. doi:10.1242/dev.090605
3.) Boudaoud, I., Fournier, É, Baguette, A., & Vallée, M. (2017). Connected Gene Communities Underlie Transcriptional Changes in Cornelia de Lange Syndrome. Genetics,207(1), 139-151. doi:10.1534/genetics.117.202291
4.) Ball, A., Chan, Y., & Yokomori, K. (2014). Mechanisms of cohesin-mediated gene regulation and lessons learned from cohesinopathies. Biochimica Et Biophysica Acta,1839(3), 191-202. doi:https://doi.org/10.1016/j.bbagrm.2013.11.002
5.) Strachan, Tom. “Cornelia De Lange Syndrome and the Link between Chromosomal Function, DNA Repair and Developmental Gene Regulation.” Current Opinion in Genetics & Development, vol. 15, no. 3, 2005, pp. 258–264., doi:10.1016/j.gde.2005.04.005.
6.) Muto, A., Calof, A. L., Lander, A. D., & Schilling, T. F. (2011). Multifactorial Origins of Heart and Gut Defects in nipbl-Deficient Zebrafish, a Model of Cornelia de Lange Syndrome [Abstract]. PLoS Biology,9(10). doi:10.1371/journal.pbio.1001181
7.) STRING Search NIPBL. (n.d.). Retrieved April 13, 2018, from https://string-db.org/
8.) Newkirk, D., Chen, Y., Chien, R., & Zeng, W. (2017). The Effect Of Nipped-B-Like (Nipbl) Haploinsufficiency On Genome-Wide Cohesin Binding And Target Gene Expression: Modeling Cornelia de Lange Syndrome. Clinical Epigenetics,9(89). doi:10.1101/134825
9.) NetPhos 3.1 Server NIPBL prediction. (n.d.). Retrieved April 15, 2018, from http://www.cbs.dtu.dk/services/NetPhos/
10.) Phosphorylation. (2018, May 09). Retrieved from https://en.wikipedia.org/wiki/Phosphorylation
Powerpoint slides :
Final talk and presentation of my specific aims.
Final talk and presentation of my specific aims.
| akers_final_presentation.pptx | |
| File Size: | 6470 kb |
| File Type: | pptx |
A condensed version of my specific aims, contains no images.
| akers_final_aims.docx | |
| File Size: | 15 kb |
| File Type: | docx |